Years ago, when the rigid licensing conditions of Mathworks, OriginLab and Wolfram and the exorbitant costs of their software began to be more than an occasional irritation, I set my hopes on the open source world. Surely there would be alternatives, I reasoned, and I would take these and be free of any annoying complications arising from proprietary software and their licensing models.
I've indeed used SciLab for a while, but eventually everyday duties and the necessity to be compatible with the rest of the world got the better of me and I returned to the commercial offers mentioned above. 😞
Now, the licensing policies of the above producers of scientific software have not, generally speaking, evolved in any desirable way—to put it mildly. It's not only the prohibitive prices, but also the involved licensing models which require considerable time to comprehend for those responsible for the purchase of this software, i.e., me.
I hence recently decided to have a closer look onto the present status of open-source scientific software. To quickly get up-to-date, I've followed discussions concerning this subject across the interwebs. The usenet proved to be particularly resourceful, and I've learned a lot from discussions there. First, I've seen that I'm not alone with my growing resentments, and second, I singled out two projects which are frequently mentioned and mostly highly commended. I'm mightily impressed by both of these projects already after the first, cursory glance.
The first of these projects is PyLab, which is perceived by many users as a viable alternative to MatLab, but is stated by its main developer to be a vision rather than a completed computational environment. The situation, however, is not nearly as bad as described in this self-critical analysis of Keir Mierle. Pylab is based on Python and the modules NumPy, SciPy, Matplotlib, and iPython, the latter being an adapted python console for interactive work. All of these modules as well as the required dependencies are readily available in modern Linux distributions. After installation, just start the console with 'ipython -pylab', and issue the following commands taken straight from the plotting tutorial:
x,y = ogrid[-1.:1.:.01, -1.:1.:.01]
z = 3*y*(3*x**2-y**2)/4 + .5*cos(6*pi * sqrt(x**2 +y**2) + arctan2(x,y))
imshow(z, origin='lower', extent=[-1,1,-1,1])
contour(z, origin='lower', extent=[-1,1,-1,1])
plot(x[:], z[50, :])
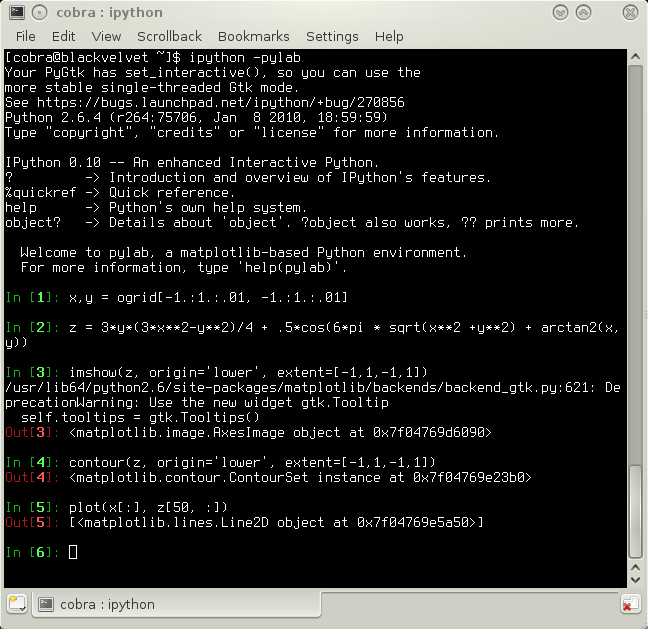
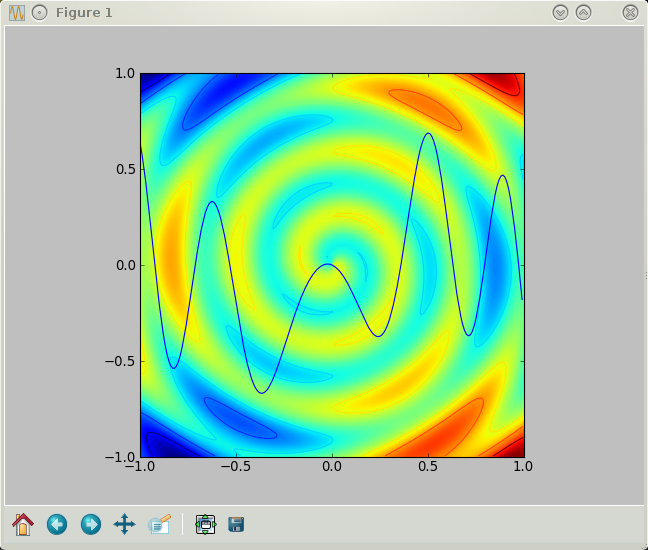
Three plots in one in five lines of code. The figure, by the way, can be saved in png format by a simple
Sympy and its isympy terminal adds symbolic abilities to pylab and is hoped to grow up to a fully-featured computer algebra system (CAS). Well, sympy is quite useful already at the present stage:

Without doubt, isympy is the most capable console calculator available. 😉
Oh, and before I forget (and since I like to clear consoles 😏):
import os
os.system('clear')
Finally, Spyder is a Qt4-based IDE for iPython and friends. As beautiful as practical!
The other project I became interested in is Sage, which is also based on python but represents an attempt to ... well, in my opinion, to become the ultimate math software on earth. 😄 Seriously, Sage is a unified interface for many major mathematical packages such as pylab, maxima, octave, scilab, R, singular, axiom, tachyon, jmol and many more I even don't know by name. Sage is executed in a client-server architecture with the client based on Ajax, thus working well in all modern web browsers.
I just see that I don't have the time to describe my first impressions of Sage in any detail anymore, since it's close to 4 pm and thus time for:
Deutschland-England
Very briefly: Sage left me speechless. Try for yourself!
It's Vuvuzela time! Yeeeehaw!